Liang Mou | Staff Software Engineer, Logging Platform Yisheng Zhou | Software Engineer II, Logging Platform Elizabeth (Vi) Nguyen | Software Engineer I, Logging Platform Owen Zhang | Senior Software Engineer, Logging Platform
Introduction
As Pinterest has grown, the demand for a robust, real-time, and cost-effective database ingestion platform has become increasingly urgent. Our data ecosystem powers a diverse set of use cases — from analytics and machine learning to product features and business intelligence — all of which depend on timely and reliable data. However, our legacy ingestion landscape was built on batch-oriented workflows and a patchwork of database dump solutions, each developed and maintained by different teams. This fragmentation made it difficult to deliver the performance, reliability, and agility required by modern data workloads.
In this blog series, we'll share our journey in building Pinterest's next-generation database ingestion framework. In this first part, we'll discuss the legacy challenges we faced, the architectural principles that shaped our new solution, and the key optimizations that enabled us to achieve significant improvements in latency, efficiency, and compliance.
Background & Motivation
The previous generation of batch-based ingestion systems presented significant challenges. First, high data latency was a critical issue, often exceeding 24 hours for updates, which severely hampered real-time analytics and machine learning applications. Second, the dependence on full-table batch jobs was inefficient. Since the daily data change for many tables was less than 5%, this approach resulted in wasted compute and storage resources by unnecessarily reprocessing unchanged records. Third, the lack of built-in support for row-level deletion created growing obstacles for maintaining data compliance. Finally, the organization suffered from operational complexity and inconsistent data quality due to the existence of multiple, independently maintained ingestion pipelines.
Our Solution: A Unified CDC-based Framework
We are introducing a unified DB ingestion framework built on Change Data Capture (Debezium/TiCDC), Kafka, Flink, Spark and Iceberg. This new system provides:
- Access to online database changes in minutes (not hours or days).
- Processing only changed records, resulting in significant infrastructure cost savings.
Native support for row-level deletion and incremental processing.
Architecture Overview
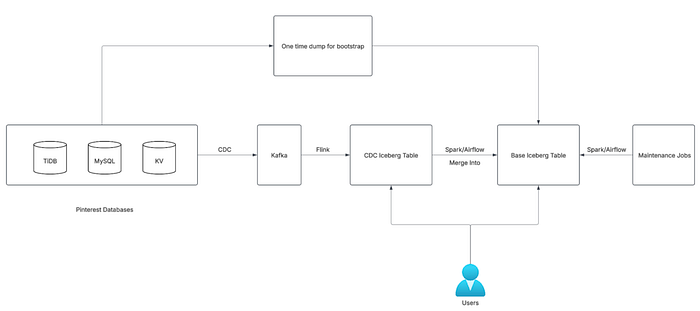
Our next-generation ingestion framework is designed to be:
- Generic: Supporting all major Pinterest databases (MySQL, KVStore, TiDB).
- Reliable: Guaranteeing at-least-once processing (zero data loss).
- Low Latency: Delivering online to offline data in minutes (typically 15 minutes to an hour).
- Scalable: Handling petabyte-scale data and thousands of pipelines.
- Config-Driven: Enabling easy onboarding via yaml config, with integrated monitoring.
Key Components
- Source Databases: The system supports Pinterest's primary databases, which are MySQL, TiDB, and KVStore.
- Change Data Capture (CDC) Layer: A custom, generic CDC service is employed to capture database changes. This service writes the events to Kafka with minimal latency, typically under one second. More details on this service are available in a previous blog post: https://medium.com/pinterest-engineering/change-data-capture-at-pinterest-7e4c357ac527.
- Streaming Layer: Flink jobs process the incoming CDC events in near real-time, persisting them into CDC Iceberg tables on S3.
- Batch Layer: Periodically (e.g., every 15 minutes), Spark jobs fetch the latest changes from the CDC tables. They then use the
Merge Intostatement to perform upserts on the Base Iceberg tables. - Bootstrap Pipeline: Before the recurring Spark upsert process begins, a dedicated bootstrap job is run to load historical data from DB dumps, initializing the Base Iceberg table.
- Maintenance Jobs: responsible for tasks like compaction and snapshot expiration.
Let's now examine in detail several crucial design choices, decisions, and optimizations.
CDC Table vs Base Table
A CDC (Change Data Capture) table functions as a time-series, append-only ledger that records every change event. The latency of the CDC table usually is under 5 minutes. An example follows:

The base table, which some refer to as a snapshot table, serves as a mirror of the online table, preserving all historical records. It maintains a direct one-to-one relationship with the upstream source table. Depending on the configuration, the latency of the base table could be 15 minutes to an hour. Here is an example:

How to update Base Table (Upsert)
The central part of this update is a two-step SparkSQL query:
- Identify Latest Updates: Process the Change Data Capture (CDC) table to retrieve the most recent update for each primary key.
- Apply Changes: Utilize a
Merge Intostatement to update the base table with these latest changes.
WITH
ranked_table AS (
SELECT
*,
ROW_NUMBER() OVER (
PARTITION BY
id
ORDER BY
ts DESC,
gtid DESC
) AS rnk
FROM
{cdc_table_name}
WHERE
dt > cast(current_date() - INTERVAL 28 DAYS as string) AND processing_ts >= to_timestamp('{previous_watermark}', 'yyyy-MM-dd HH:mm:ss.SSS')
),
deduped_ranked_table AS (
SELECT * from ranked_table where rnk = 1
)
MERGE INTO
{base_table_name} T USING (
SELECT
id,
data.name,
data.email,
data.address
FROM
deduped_ranked_table
) s ON t.id = s.id
WHEN MATCHED AND s.type = 'delete' THEN DELETE
WHEN MATCHED THEN UPDATE SET
t.name = s.name,
t.email = s.email,
t.address = s.address,
WHEN NOT MATCHED AND s.type != 'delete' THEN INSERT *;Iceberg's Merge Intooperation offers two distinct strategies, each with its own set of tradeoffs. After evaluating the two strategies at Pinterest, we've standardized on the Merge-on-Read (MOR) approach. The Copy-on-Write (COW) strategy was deemed unsuitable for most of our use cases because its significantly higher storage cost outweighs its other potential benefits.
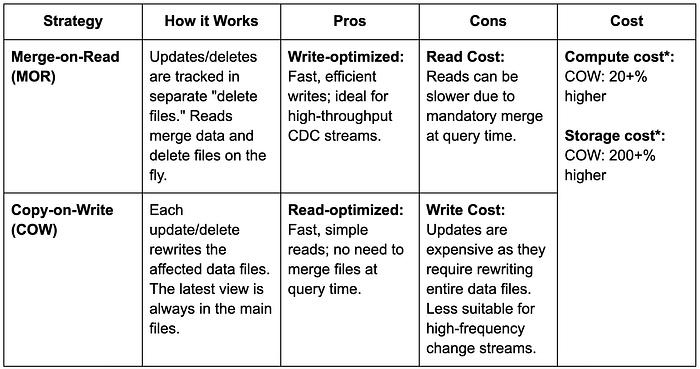
* Compute Cost: COW incurs higher computational costs due to the increased overhead of write operations.
* Storage Cost: COW's storage expenses are significantly elevated because the new data files generated with each snapshot are substantially larger.
Optimizations
We've identified and resolved numerous bottlenecks during this process, implementing various optimizations to enhance the efficiency of the Upsert operation. Below are a few examples.
Partitioning the Base Table by Primary Key Hash
For larger tables, partitioning the base table by a hash of the primary key (using the bucket function) is generally recommended. This approach distributes records evenly across partitions, which can significantly improve the performance of Upsert operations. When Spark processes an upsert, it can operate on each partition in parallel, reducing the amount of data that needs to be scanned and rewritten. For example, partitioning by bucket(100, id) allows Spark and Iceberg to parallelize updates across 100 partitions, making the upsert process much more efficient. Below is an example of how to enable bucketing.
CREATE TABLE IF NOT EXISTS {base_table_name} (
id BIGINT,
name STRING,
email STRING,
address STRING
)
USING iceberg
LOCATION 's3://xxx'
PARTITIONED BY (bucket(100, id))
TBLPROPERTIES(
'format-version' = '2',
'identifier-fields' = '[id]',
'write.upsert.enabled' = 'true',
'write.delete.mode' = 'merge-on-read',
'write.merge.mode' = 'merge-on-read',
'write.update.mode' = 'merge-on-read',
'read.split.target-size' = '1073741824',
'write.target-file-size-bytes' = '1073741824'
);Small files problem
After introducing bucketing, we noticed that each upsert operation was generating a large number of small files within each partition. Upon investigation, we discovered that during upserts, each Spark task could write files for multiple partitions, which led to an explosion of small files — especially in larger clusters with many parallel tasks.
To address this, we set the table property to WRITE DISTRIBUTED BY PARTITION. This change instructs Spark (and Iceberg) to organize the write workload so that all data for a given partition is grouped and written together. As a result, this significantly reduces the number of small files per partition and improves overall table performance.
ALTER TABLE {base_table_name} WRITE DISTRIBUTED BY PARTITION;Bucket join for large tables
Working with extremely large base and CDC (Change Data Capture) tables revealed that standard MERGE INTO operations, which rely on joins, were prohibitively expensive due to Spark's default behavior of potentially shuffling the entire base table for alignment.
To solve this, we implemented Bucket Join. While Bucket Join typically requires the CDC and Base tables to have identical partitioning schemas, and we prefer to partition the CDC table by time (dt/hr) rather than the Primary Key (PK) for other use cases, we developed a workaround. We introduce a temporary table, populate it with the CDC data, and then use this temporary table to perform the Upsert against the base table.
This approach allows Spark to match corresponding data buckets directly, completely bypassing a full shuffle of the massive base table before each merge operation. Our benchmarks demonstrated significant efficiency gains, resulting in a 40%+ reduction in compute cost as well as significantly reduced latency.
CREATE TABLE IF NOT EXISTS {cdc_bucketed_table_name} (
id BIGINT,
name STRING,
email STRING,
address STRING
)
USING iceberg
LOCATION 's3://xxx'
PARTITIONED BY (bucket(100, id));
WITH
ranked_table AS (
SELECT
*,
ROW_NUMBER() OVER (
PARTITION BY
id
ORDER BY
ts DESC,
gtid DESC
) AS rnk
FROM
{cdc_table_name}
WHERE
dt > cast(current_date() - INTERVAL 28 DAYS as string) AND processing_ts >= to_timestamp('{previous_watermark}', 'yyyy-MM-dd HH:mm:ss.SSS')
),
deduped_ranked_table AS (
SELECT * from ranked_table where rnk = 1
)
INSERT OVERWRITE TABLE {cdc_bucketed_table_name}
SELECT
id,
data.name,
data.email,
data.address
FROM
deduped_ranked_table;
MERGE INTO
{base_table_name} T USING (
SELECT
id,
name,
email,
address
FROM
{cdc_bucketed_table_name}
) s ON t.id = s.id
WHEN MATCHED AND s.type = 'delete' THEN DELETE
WHEN MATCHED THEN UPDATE SET
t.name = s.name,
t.email = s.email,
t.address = s.address,
WHEN NOT MATCHED AND s.type != 'delete' THEN INSERT *;Conclusion & What's Next
In this post, we've walked through the motivations behind building a unified, real-time database ingestion framework at Pinterest, the architectural choices that shaped our solution, and the key optimizations that enabled us to deliver low-latency, cost-efficient, and reliable data pipelines at scale. By leveraging Change Data Capture, modern data lake technologies, and a series of targeted performance improvements, we've been able to overcome the limitations of legacy batch workflows and fragmented pipelines.
But our journey doesn't stop here. One of the most challenging aspects of building a robust ingestion platform is supporting seamless, automated schema evolution — ensuring that changes in upstream databases are safely and efficiently reflected downstream, without breaking data consumers or requiring manual intervention.
In the next installment of this blog series, we'll dive deep into how we designed and implemented automated schema evolution in our CDC-based ingestion framework. We'll share the technical hurdles we faced, the solutions we developed, and the lessons we learned along the way.
Stay tuned for Part 2: Automated Schema Evolution!
Thank you for reading, and we look forward to sharing more soon.
Acknowledgments
The success of this project would not have been possible without the significant contributions and support of:
- Ads database infra: Elly Chen, Jackie Xu, Chengcheng Hu
- Ads indexing infra: Bingxin Zhang, Fredy Virguez, Yuhan Peng
- Ads retrieve infra: Zhicheng Jin, Hugo Milhomens, Tao Yang
- Storage services: Gabriel Raphael Garcia Montoya, Yuan Gao, Leonardo Marques Maciel Silva, Liqi Yi, Alberto Ordonez Pereira, Lianghong Xu
- Storage foundations: Tailin Lyu, Yu Su, John Grass, Alberto Ordonez Pereira, Lianghong Xu
- Streaming/Batch processing: Kanchi Masalia, Kevin Browne, Tucker Harvey, Chen Qin
- Big data storage: Pucheng Yang, Mian Luo, Roman Horilyi
- Big data processing: Zaheen Aziz
- Big data infra: Zheyu Zha
- Big data: Yi Pan, Ashish Singh
- Data privacy: Keith Regier
- Github: Nathan Hilton
- Logging team: Vahid Hashemian, Jeff Xiang, Artem Tetenkin
Special gratitude must be extended to Sharddul Jewalikar, Ang Zhang and Roger Wang for their continuous guidance, feedback, and support throughout the project.
Disclaimer
Apache®️, Apache Flink®️, Apache Iceberg®️, Apache Kafka®️, Apache Spark®️,and Kafka®️ are trademarks of the Apache Software Foundation (https://www.apache.org/).
Amazon®️, AWS®️, S3®️, and EC2®️ are trademarks of Amazon.com, Inc. or its affiliates.
Debezium®️ is a trademark of Red Hat, Inc.
MySQL®️ is a trademark of Oracle Corporation.
RocksDB®️ is a trademark of Meta Platforms, Inc. or its affiliates.
TiDB®️ is a trademarks of Beijing PingCAP Xingchen Technology and Development Co.


