Pandas is powerful — but CPU-hungry. Here's how I profiled and sped up my Pandas workflows with smarter memory use and parallel execution.
🧠 The Illusion of Simplicity
Pandas is every data analyst's comfort zone. With a few lines of code, you can read a CSV, clean the data, and run analysis.
But here's what I learned the hard way:
Pandas is deceptively slow at scale.
I was feeding it millions of rows and wondering why my machine was melting. Turns out, Pandas will gladly eat your CPU alive unless you tame it.
🧪 The First Sign: My Laptop Fan Screamed Like a Jet Engine
I loaded a 5GB CSV file and started chaining operations like a maniac:
df = pd.read_csv('big.csv')
df = df[df['active'] == True]
df['score'] = df['clicks'] / df['views']
df = df.sort_values('score', ascending=False)Everything worked… but it took forever.
💻 CPU: Maxed 📈 Memory: Spiked ⏳ Me: Waiting 3+ minutes per run
That's when I knew: Pandas wasn't the problem — I was.
🕵️ Step 1: Profile Before You Optimize
Before reaching for parallel tools or rewriting code, I used:
import pandas as pd
import time
start = time.time()
# your pandas workflow
end = time.time()
print(f"Execution time: {end - start:.2f}s")Or for granular timing:
%timeit df['score'] = df['clicks'] / df['views']For memory:
import sys
df.memory_usage(deep=True).sum() / 1024**2 # In MBThen came the real game-changer:
pip install line_profilerProfile exact lines and know where time was leaking.
⚙️ Step 2: Eliminate the Slowest Offenders
🔄 1. Chained Operations
Bad:
df = df[df['active'] == True].sort_values('score').drop_duplicates()Better:
df = df[df['active'] == True]
df = df.sort_values('score')
df = df.drop_duplicates()✅ Easier to profile ✅ Easier to parallelize later
📊 2. Unvectorized Loops
This is a CPU killer:
for i in range(len(df)):
df.loc[i, 'adjusted'] = df.loc[i, 'score'] * 1.1Use vectorization instead:
df['adjusted'] = df['score'] * 1.1🚀 Step 3: Parallelize What Pandas Can't
Pandas is single-threaded. So for big datasets, even optimized code chokes.
That's where modular parallelism shines.
Option 1: Use swifter (drop-in parallelization)
pip install swifterThen:
import swifter
df['score'] = df.swifter.apply(my_func, axis=1)It auto-detects when to use Dask or vectorized ops.
Option 2: Chunk Processing
Split big files:
chunks = pd.read_csv('big.csv', chunksize=100_000)
for chunk in chunks:
process(chunk)Memory-efficient and fast.
Option 3: Dask — Parallel Pandas
pip install dask
import dask.dataframe as dd
ddf = dd.read_csv('big.csv')
ddf = ddf[ddf['active'] == True]
result = ddf.compute()Uses all CPU cores. Almost the same syntax as Pandas.
🧼 Step 4: Clean Up the Memory Hog
🔍 Use category for low-cardinality columns
df['country'] = df['country'].astype('category')🧹 Remove unused columns
df.drop(['col1', 'col2'], axis=1, inplace=True)🧊 Force data types early
df = pd.read_csv('big.csv', dtype={
'user_id': 'int32',
'score': 'float32'
})Saves gigabytes when done right.
⚡ Final Checklist: From CPU Hog to Speed Demon
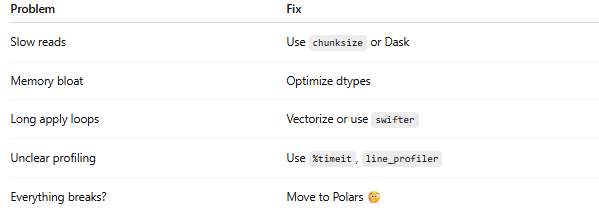
🧠 Final Thoughts
Pandas is fantastic — but not magic.
It was never meant to scale to hundreds of millions of rows, but it can if you respect the CPU and optimize intentionally.
So the next time your fan screams, remember:
Treat Pandas like a CPU hog. Feed it wisely — or watch it eat your machine alive.
