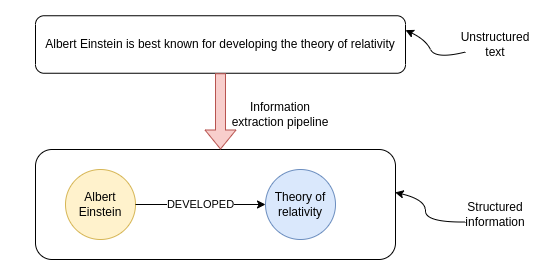
Since I first dabbled with natural language processing, I have had a special place in my heart for information extraction (IE) pipelines. Information extraction (IE) pipelines extract structured data from unstructured data like text. The internet provides an abundance of information in the form of various articles and other content formats. However, while you might read the news or subscribe to multiple podcasts, it is virtually impossible to keep track of all the new information released daily. Even if you could manually read all the latest reports and articles, it would be incredibly tedious and labor-intensive to structure the data so that you can easily query and aggregate it with your preferred tools. I definitely wouldn't want to be doing that as my job. Luckily, we can resort to using the latest state-of-the-art NLP techniques to do the information extraction for us automatically.
While I have already implemented and written about an IE pipeline, I've noticed many new advancements in open-source NLP models, particularly around spaCy. I later learned that most of the models I will be using in this post are simply wrapped as a spaCy component, and you could use other libraries if you liked. However, since spaCy was the first NLP library I've played around with, I've decided to implement the IE pipeline in spaCy as a way of saying thanks to the developers for making such a great and easy to get started tool.
How I perceive the steps in the IE pipeline has remained identical over time.
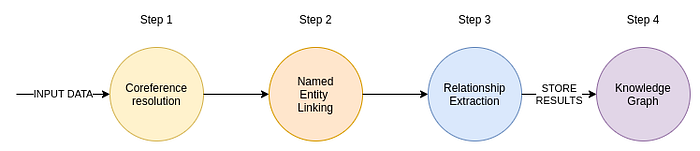
The input to the IE pipeline is text. That text can come from articles or perhaps internal business documents. If you deal with PDFs or images, you could use computer vision to extract the text. Also, we could use voice2text models to convert audio recordings into text. After preparing the input text, we first run it through the coreference resolution model. The coreference resolution converts pronouns into referred entities. Most often, the examples for coreference resolution are personal pronouns, where, for example, the model replaces pronouns with the referred person's name.
The next step is identifying entities in the text. Which entities we want to recognize is entirely up to the use case we are dealing with and can vary from domain to domain. For example, you will often see NLP models trained to identify people, organizations, and locations. However, in the biomedical domain, we may want to identify concepts like genes, diseases, etc. I've also seen some examples where internal documents of a business are being processed in order to construct a knowledge base that could point to documents that contain answers a user might have or even fuel a chatbot. The process of identifying entities in the text is known as named entity recognition.
Identifying relevant entities in the text is one step, but you almost always need to standardize entities. For example, say that the text references the Theory of relativity and Relativity theory. It might seem obvious that the two entities refer to the same concept. However, as much as it is simple for you, we want to avoid manual labor as much as possible. And a machine does not automatically recognize both references as the same concept. Here is where the named entity disambiguation or entity linking comes into play. The goal of both the named entity disambiguation and entity linking is to assign a unique id to all the entities in the text. With entity linking, extracted entities from the text are mapped to corresponding unique ids from a target knowledge base. In practice, you will see Wikipedia being used as the target knowledge base a lot. However, if you are working in more specific domains or want to process internal business documents, perhaps Wikipedia might not be the best target knowledge base. Remember, an entity must exist in the target knowledge base for the entity linking process to be able to map entities from text to it. If the text mentions you and your manager, and both of you are not on Wikipedia, then it doesn't make sense to use Wikipedia as the target knowledge base in the entity linking process.
Lastly, the IE pipeline then uses relation extraction models to identify any relationships between text that are mentioned in the text. If we hold on to the manager example, let's say we have the following text.
Alicia is the manager of Zach.
Ideally, we would want the relation extraction model to recognize the relationship between the two entities.
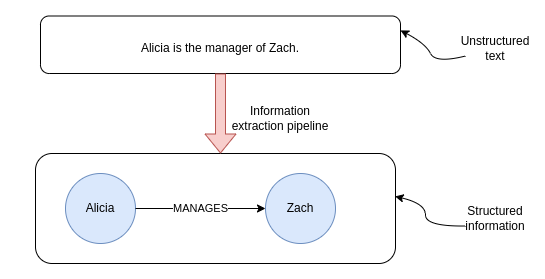
Most of relation extraction models are trained to recognize multiple types of relationships, so that the information extraction is as rich as possible.
Now that we quickly went over the theory, we can jump to the practical examples.
Developing IE pipeline in spaCy
There has been much development around spaCy in the last couple of weeks, so I decided to try out the new plugins and use them to construct an information extraction pipeline.
As always, all the code is available on Github.
Coreference resolution
First off, we are going to be using the new Crosslingual Coreference model contributed by David Berenstein to the spaCy Universe. SpaCy Universe is a collection of open-source plugins or addons for spaCy. The cool thing about the spaCy universe project is that it's straightforward to add the models to our pipeline.
That's it. It only took a couple of lines to set up the coreference model in spaCy. We can now test out the coreference pipeline.
As you can notice, the personal pronoun He was replaced with the referred person Christian Drosten. As simple as it seems, this is a vital step to developing an accurate information extraction pipeline. Massive shoutout to David Berenstein for continually developing this project and making our life easier.
Relation extraction
You might wonder why we skipped the named entity recognition and linking step. Well, the reason is that we will be using the Rebel project that recognizes both the entities and relations from the text. If I understand correctly, the Rebel project was developed by Pere-Lluís Huguet Cabot as a part of his PhD study with Babelscape and Sapienza University. Again, a massive shoutout to Pere for creating such an incredible library with state-of-the-art results for relation extraction. The Rebel model is available on Hugginface as well as in the form of spaCy component.
However, the model doesn't do any entity linking, so we will implement our version of entity linking. We will simply search for entities on WikiData by calling the search entities WikiData API.
As you can notice from the code, we simply take the entity id from the first result. I've been looking at how to improve this and stumbled upon the ExtEnd project. ExtEnd project is a novel approach to entity disambiguation and is available as a demo on Huggingface as well as a spaCy component. I've played around with it a bit and managed to get it working using WikiData API for candidates instead of the original AIDA candidates. However, when I wanted to have all three projects (Coref, Rebel, ExtEnd) in the same pipeline, there were some dependency issues as they use a different version of PyTorch, so I gave up for now. I guess I could dockerize the pipeline to solve the dependency issues, but I wanted to have both the ExtEnd and the Rebel in a single spaCy pipeline. However, the ExtEnd code I developed is available on GitHub, and if someone wants to help me get it working, I am more than happy to accept Pull requests.
Ok, so for now, we won't be using the ExtEnd project but will use a simplified version of entity linking by simply taking the first candidate fetched from the WikiData API. The only thing we need to do is incorporate our simplified entity linking solution in the Rebel pipeline. Since the Rebel component is not available directly as a spaCy Universe project, we must copy the component definition from their repository manually. I've taken the liberty to implement my version of the set_annotations function in the Rebel spaCy component, while the rest of the code is the same as the original.
The set_annotations function handles how we store the results back to the spaCy's Doc object. First, we ignore all the self-loops. Self-loops are relationships that start and end at the same entity. Next, we search for both the head and tail entities of the relation in the text using regex. I've noticed that the Rebel model sometimes hallucinates some entities which are not in the original text. For that reason, I added a step that verifies that both entities are actually in the text before appending them to the results.
Lastly, we use the WikiData API to map extracted entities to WikiData ids. As mentioned, this is a simplified version of entity disambiguation and linking, and you can take a more novel approach like the ExtEnd model, for example.
Now that the Rebel spaCy component is defined, we can create a new spaCy pipeline to handle the relation extraction part.
Finally, we can test the relation extraction pipeline on the sample text we've used before for coreference resolution.
The Rebel model extracted two relations from the text. For example, it recognized that Christian Drosten, with the WikiData id Q1079331, is employed by Google, which has an id Q95.
Storing the information extraction pipeline results
Whenever I hear about relation information between entities, I think of a graph. A graph database is developed to store relations between entities, so what better fit to store the information extraction pipeline results.
As you might know, I am biased towards Neo4j, but you can use whatever tool you like. Here, I will demonstrate how to store the results of my implementation of the information extraction pipeline into Neo4j. We will process a couple of Wikipedia summaries of famous women scientists and store the results as a graph.
If you want to follow with the code examples, I would suggest you to create a Blank Project in Neo4j Sandbox environment. After you have created the Neo4j Sandbox instance, you can copy the credentials to the code.
Next, we need to define the function that will retrieve Wikipedia summaries for the famous women scientists, run the text through the information extraction pipeline, and finally store the results to Neo4j.
We've used the wikipedia python library to help us fetch the summaries from Wikipedia. Next, we need to define the Cypher statement used to import the information extraction results. I won't go into details of Cypher syntax, but basically, we first merge the head and tail entities by their WikiData id and then use a procedure from the APOC library to merge the relationship. I recommend going through courses in the Neo4j Graph Academy if you are looking for resources to learn more about Cypher syntax.
Now that we have everything ready, we can go ahead and parse a couple of Wikipedia summaries.
Once the processing is finished, you can open Neo4j Browser to inspect the results.
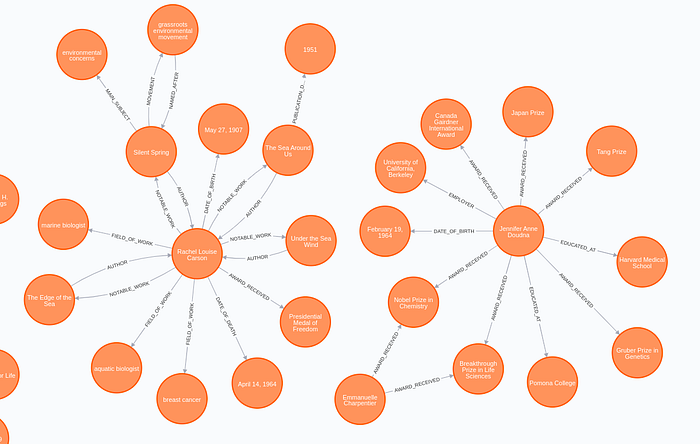
The results look surprisingly well. In this example, the Rebel model recognized more than 20 relation types between entities ranging from AWARD and EMPLOYER to DRUG_USED_FOR_TREATMENT. Just for fun, I'll show you the biomedical relation the model extracted.
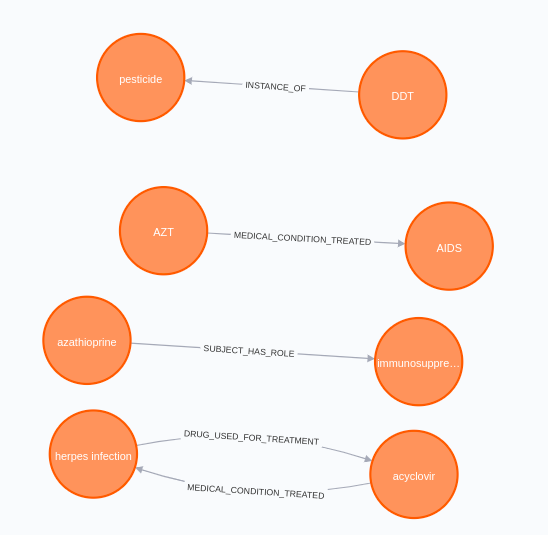
It seems that some of those ladies worked in the biomedical domain. Interestingly, the model identified that acyclovir is used to treat herpes infection, and that azathioprine is an immunosuppressive drug.
Enriching the graph
Since we have mapped our entities to the WikiData ids, we can further use the WikiData API to enrich our graph. I will show you how to extract INSTANCE_OF relations from WikiData and store them to Neo4j with the help of the APOC library, which allows us to call web APIs and store results in the database.
To be able to call WikiData API, you need to have a basic understanding of SPARQL syntax, but that is beyond the scope of this blog post. However, I've written a post some time about that shows more SPARQL queries used to enrich a Neo4j graph and delves into SPARQL syntax.
By executing the following query, we add the Class nodes to the graph and link them with the appropriate entities.
Now we can inspect the results of the enrichment in Neo4j Browser.
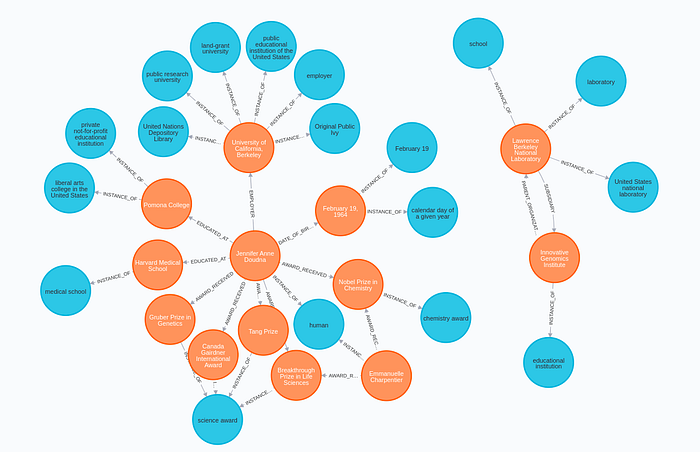
Conclusion
I really like what spaCy is doing lately and all the open-source projects around it. I noticed that various open-source projects are primarily standalone, and it can be tricky to combine multiple models into a single spaCy pipeline. For example, you can see that we had to have two pipelines in this project, one for coreference resolution and one for relation extraction and entity linking.
As for the results of the IE pipeline, I am delighted with how well it turned out. As you can observe in the Rebel repository, their solution is state-of-the-art on many NLP datasets, so it is not a big surprise that the results are so good. The only weak link in my implementation is the Entity Linking step. As I said, it would probably greatly benefit from adding something like the ExtEnd library for more accurate entity disambiguation and linking. Perhaps that's something I'll do next time.
Try out the IE implementation, and please let me know what you think or if you have some ideas for improvements. There are a ton of opportunities to make this pipeline better!
As always, the code is available on GitHub.